What does a low 260 280 ratio tell you about your DNA sample
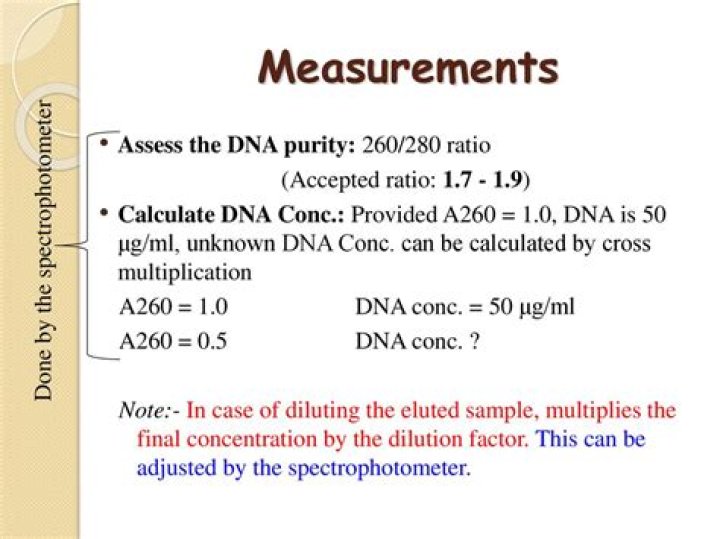

A 260/280 ratio of ~1.8 is generally accepted as “pure” for DNA; a ratio of ~2.0 is generally accepted as “pure” for RNA. Abnormal 260/280 ratios usually indicate that a sample is contaminated by residual phenol, guanidine, or other reagent used in the extraction protocol, in which case the ratio is normally low.
What does a low 260 280 ratio indicate?
260/280 ratio – a low ratio may be the result of a contaminant absorbing at 280 nm or less. Wavelength of the trough in sample spectrum– this should be at ~230 nm. Absorbance by a contaminant at a low wavelength will typically shift the wavelength of the trough.
Why is it important to interpret the 260 nm 280 nm ratios from the result of spectrophotometer?
The ratio of absorbance at 260 nm and 280 nm is used to assess the purity of DNA and RNA. … If the ratio is appreciably lower in either case, it may indi- cate the presence of protein, phenol or other contaminants that absorb strongly at or near 280 nm.
What is an acceptable 260 280 ratio for DNA?
The ratio of absorbance at 260 and 280 nm is used to assess DNA purity. A ratio of ∼1.8 is generally accepted as “pure” for DNA. If the ratio is appreciably lower (≤1.6), it may indicate the presence of proteins, phenol, or other contaminants that absorb strongly at or near 280 nm.What does a low 260 230 mean?
• 260/230 ratio – a low ratio may be the result of a contaminant absorbing at 230 nm or less. • 260/280 ratio – a low ratio may be the result of a contaminant absorbing at 280 nm or less.
What is a good DNA concentration ng uL?
for DNA sizes above 500bp, it is recommended the minimum concentration is 40ng/ul with a minimum volume of 15uL. for sizes below 500bp, 20ng/uL is sufficient.
What is a good DNA concentration?
The most common purity calculation is the ratio of the absorbance at 260nm divided by the reading at 280nm. Good-quality DNA will have an A260/A280 ratio of 1.7–2.0. A reading of 1.6 does not render the DNA unsuitable for any application, but lower ratios indicate more contaminants are present.
Why does DNA absorb at 260?
Nucleic acids strongly absorb UV light with wavelengths of 260 nm due to the resonance structure of the purine and pyrimidine bases [7]. The absorbance is converted into ng/μL of double stranded DNA (dsDNA) using the established conversion factor of 50 ng/μL for 1 optical density unit at 260 nm [9].What does a 260 280 ratio above 2 mean?
It is a sign of RNA contamination. … 260/ 280 ratio of ~1.8 is generally accepted as “pure” for DNA and a ratio of ~2.0 is generally accepted as “pure” for RNA. For any DNA sample with A 260/280 ratio more than 1.8 indicates the presence of RNA as contamination.
What absorbs at 280nm?Specifically, the amino acids tyrosine and tryptophan have a very specific absorption at 280 nm, allowing direct A280 measurement of protein concentration. UV absorbance at 280 nm is routinely used to estimate protein concentration in laboratories due to its simplicity, ease of use and affordability.
Article first time published onWhat is a good 260 280 ratio for protein?
An ideal 260/280 ratio for common proteins is 0.6. Higher ratios may indicate the contamination of isolated proteins with DNA. Alternatively, the buffer used to isolate the sample protein may include components that absorb strongly in the UV region.
What does A280 mean?
Protein has a high absorbance at 280 while nucleic acids have a high absorbance at 260. On the other hand contaminants like phenol have a high absorbance at 230. High A230 or A280 indicates that your nucleic acid prep has a high amount of contamination which may or may not affect downstream assays.
Why do proteins absorb light at 280 nm?
Proteins in solution absorb ultraviolet light with absorbance maxima at 280 and 200 nm. Amino acids with aromatic rings are the primary reason for the absorbance peak at 280 nm. Peptide bonds are primarily responsible for the peak at 200 nm.
What absorbs at 230nm?
Absorbance at 230 nm Many organic compounds have strong absorbances at around 225 nm. In addition to phenol, TRIzol, and chaotropic salts, the peptide bonds in proteins absorb light between 200 and 230 nm.
How do you increase RNA 260 230?
I usually improve my 260/230 ratios by doing a re-precipitation with sodium acetate / ethanol. If you get some precipitates or gunk, try to dissolve them as best as you can after adding the sodium acetate, then vigorously vortex again after adding ethanol (3x10s).
What is a good DNA concentration NanoDrop?
If using a NanoDrop to measure your samples, place 1-2µL of mini-prepped DNA onto the pedestal. … Note: Purity is measured under the 260/280 column (A good purity ranges from 1.80-2.00).
What is the best DNA concentration for PCR?
50-100ng genomic DNA and aprox. 20-50ng of plasmid is sufficient for PCR.
What is a good DNA concentration for PCR?
Normally used concentration are 100-250 ng for mammalian genomic DNA and 20 ng for linearized plasmid DNA (circular plasmid DNA is slightly less efficiently amplified) per 50µl reaction.
Why is DNA concentration low?
The most common causes for low yield are poor culturing conditions and plasmid propagation, excessive amounts of starting material resulting in insufficient bacterial cell lysis and column overloading.
What is the minimum amount of DNA needed for PCR?
For PCR reactions, we recommend using 5-10ng per PCR reaction and at least 2ng genomic DNA should be included in each PCR reaction.
How can I increase my DNA concentration?
- Add 1/10 volume of 3 M sodium acetate (pH = 5.2) to the sample in solution.
- Add 1 uL of glycogen solution (20 mg/mL) per 20 μL of sample in solution.
- Add 1.0 volume of isopropanol.
- Incubate at -20° C for 1 hour.
- Centrifuge 15 minutes at 10,000 rpm.
What is the significance of two wavelength 260 nm and 280 nm in DNA quantification?
It is based on the principles that nucleic acids absorb ultraviolet (UV) light at a specific wavelength. For pure DNA samples, the maximum absorbance occurs over a broad peak at around 260 nm; at 280 nm it only absorbs about half as much UV light compared to 260 nm [2].
How is DNA RNA contamination detected?
RNA samples need to be DNA-free. The RNA isolation protocol should always include a DNase digestion step; in problematic cases use RNA-clean & concentrator kits with DNase. On an agarose gel, DNA contamination will be visible as a smear or band of fragments considerably larger than the RNA (>10 kb).
What does the ratio A260 A280 Tell us about the purity of a sample of purified DNA?
The A260/A280 provides insight regarding the type of nucleic acid present (dsDNA or RNA) as well as providing a rough indication of purity. Typically, protein contamination can be detected by a reduction of this ratio; RNA contamination can be detected by an increase of this ratio.
Which of the following A260 280 ratios would be a reasonable cut off for DNA that you intend to use for PCR experiments?
Which of the following A260/280 ratios would be a reasonable cut off for RNA that you intend to use for quantitative RT-PCR? Pure RNA is generally accepted as having an A260/280 ratio of 2.0.
Do proteins absorb at 260 nm?
Nucleic acids absorb light at 260 nm and proteins absorb at 280 nm. Therefore, a high value indicates the presence of more nucleic acids and a low value indicates the presence of proteins.
Does 280 absorb ATP?
when measuring the protein concentration we routinely use BioRad’s Bradford assay because ATP absorbs at 280nm.
Do all proteins absorb at 280?
Summary. Proteins absorb strongly at 280 nm due to three types of its constituent amino acids. The peptide bonds found in the amino acids also absorb at 205 nm. The UV absorption of protein can be used both to quickly image and acquire spectra of microscopic samples non-destructively.
Which amino acids absorb UV light at 280?
Tyrosine and tryptophan absorb more than do phenylalanine; tryptophan is responsible for most of the absorbance of ultraviolet light (ca. 280 nm) by proteins. Tyrosine is the only one of the aromatic amino acids with an ionizable side chain. Tyrosine is one of three hydroxyl containing amino acids.
What chromophores are responsible to UV absorption in a protein?
The chromophore responsible for light absorption in PYP is a p-coumaric acid (see Fig. 1 A). In the ground state the chromophore is in the trans form, the phenol group is deprotonated (4,5), and it is covalently linked to the protein via a thioester linkage to cysteine residue 69.
What is a good A280?
For a pure protein, the A260/A280 ratio should be 0.5-0.55; higher values suggest nucleic acid contamination. Nucleic acids will also lead to an overestimation of the protein concentration. … The best would be to produce a pure, structured protein and make a standard curve, if this is possible!